What did scientists discover?
Intact protein analysis ("top down proteomics") by mass spectrometry attempts to fully characterize proteins found in biological systems. This can be a difficult problem, as even a relatively simple extract from bacteria contains thousands of proteins.
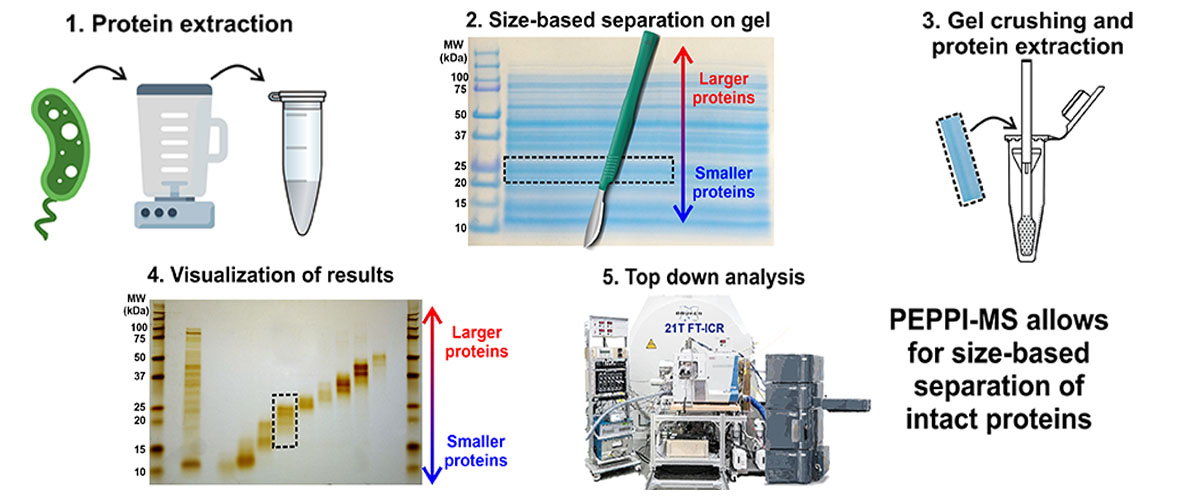
The complex mixture of proteins can be simplified by prefractionation, in which protein mixtures are separated into several simpler fractions based on size. Together with collaborators at Ehime University in Japan, a new method for prefractionation was developed called PEPPI-MS (Passively Eluting Proteins from Polyacrylamide gels as Intact species for Mass Spectrometry). This new method is much simpler and less expensive than the previous state-of-the-art for this rapidly growing research frontier.
In recognition of the importance of this new technique, the PEPPI-MS publication was selected ACS Editors' Choice® and granted free public access.
Why is this important?
Existing prefractionation methods require expensive, dedicated equipment, which limits access to only a few researchers. PEPPI-MS uses low-cost materials and equipment commonly found in biochemistry laboratories, which makes an important separation strategy for top down proteomics widely available.
Who did the research?
David S. Butcher1, Ayako Takemori2, Nobuaki Takemori2, and Lissa C. Anderson1
1National MagLab, Florida State University, Tallahassee, FL 2Advanced Research Support Center, Ehime University, Toon, Ehime, Japan
Why did they need the MagLab?
Development and optimization of the PEPPI-MS protein sample preparation technique required advanced characterization of protein mixtures using the MagLab's unique 21T FT-ICR mass spectrometry instrument.
Details for scientists
- View or download the expert-level Science Highlight, Prefractionation of Intact Proteins for Mass Spectrometry
- Read the full-length publication, PEPPI-MS: Polyacrylamide-Gel-Based Prefractionation for Analysis of Intact Proteoforms and Protein Complexes by Mass Spectrometry, in J. Proteome Research
Funding
This research was funded by the following grants: Funding Grants: G.S. Boebinger (NSF DMR-1644779); N. Takemori (JSPS KAKHENI 16K08937, 18H04559, 19K05526
For more information, contact Chris Hendrickson.